Compute mean, sd, and CV for all Peptides, or proteins, for all interactions and all samples.
Source:R/tidyMS_stats.R
summarize_stats.RdCompute mean, sd, and CV for all Peptides, or proteins, for all interactions and all samples.
Compute mean, sd, and CV for e.g. Peptides, or proteins, for all samples.
summarize stats output (compute quantiles)
See also
Other stats:
INTERNAL_FUNCTIONS_BY_FAMILY,
lfq_power_t_test_proteins(),
lfq_power_t_test_quantiles(),
lfq_power_t_test_quantiles_V2(),
plot_stat_density(),
plot_stat_density_median(),
plot_stat_violin(),
plot_stat_violin_median(),
plot_stdv_vs_mean(),
pooled_V2()
Other stats:
INTERNAL_FUNCTIONS_BY_FAMILY,
lfq_power_t_test_proteins(),
lfq_power_t_test_quantiles(),
lfq_power_t_test_quantiles_V2(),
plot_stat_density(),
plot_stat_density_median(),
plot_stat_violin(),
plot_stat_violin_median(),
plot_stdv_vs_mean(),
pooled_V2()
Other stats:
INTERNAL_FUNCTIONS_BY_FAMILY,
lfq_power_t_test_proteins(),
lfq_power_t_test_quantiles(),
lfq_power_t_test_quantiles_V2(),
plot_stat_density(),
plot_stat_density_median(),
plot_stat_violin(),
plot_stat_violin_median(),
plot_stdv_vs_mean(),
pooled_V2()
Examples
bb <- prolfqua::sim_lfq_data_protein_config()
#> creating sampleName from file_name column
#> completing cases
#> completing cases done
#> setup done
lfq <- LFQData$new(bb$data, bb$config)
res1 <- summarize_stats(lfq)
res2 <- prolfqua::sim_lfq_data_2factor_config()
#> creating sampleName from file_name column
#> completing cases
#> completing cases done
#> setup done
res2$config$factor_depth <- 2
lfq2 <- LFQData$new(res2$data, res2$config)
stats <- summarize_stats(lfq2)
stopifnot(nrow(stats) == 40)
stats <- summarize_stats(lfq2, factor_key = lfq2$factor_keys()[1])
stopifnot(nrow(stats) == 20)
stats <- summarize_stats(lfq2, factor_key = lfq2$factor_keys()[2])
stopifnot(nrow(stats) == 20)
stats <- summarize_stats(lfq2, factor_key = NULL)
stopifnot(nrow(stats) == 10)
bb <- prolfqua::sim_lfq_data_protein_config()
#> creating sampleName from file_name column
#> completing cases
#> completing cases done
#> setup done
lfq <- LFQData$new(bb$data, bb$config)
res1 <- summarize_stats_all(lfq)
stopifnot((res1 |> dplyr::filter(group_ == "All") |> nrow()) == (res1 |> nrow()))
library(ggplot2)
bb1 <- prolfqua::sim_lfq_data_peptide_config()
#> creating sampleName from file_name column
#> completing cases
#> completing cases done
#> setup done
lfq <- LFQData$new(bb1$data, bb1$config)
stats_res <- summarize_stats(lfq)
sq <- summarize_stats_quantiles(stats_res, lfq$relevant_factor_keys())
sq <- summarize_stats_quantiles(stats_res, lfq$relevant_factor_keys(), stats = "CV")
sq <- summarize_stats_quantiles(stats_res, lfq$relevant_factor_keys(), stats = "sd")
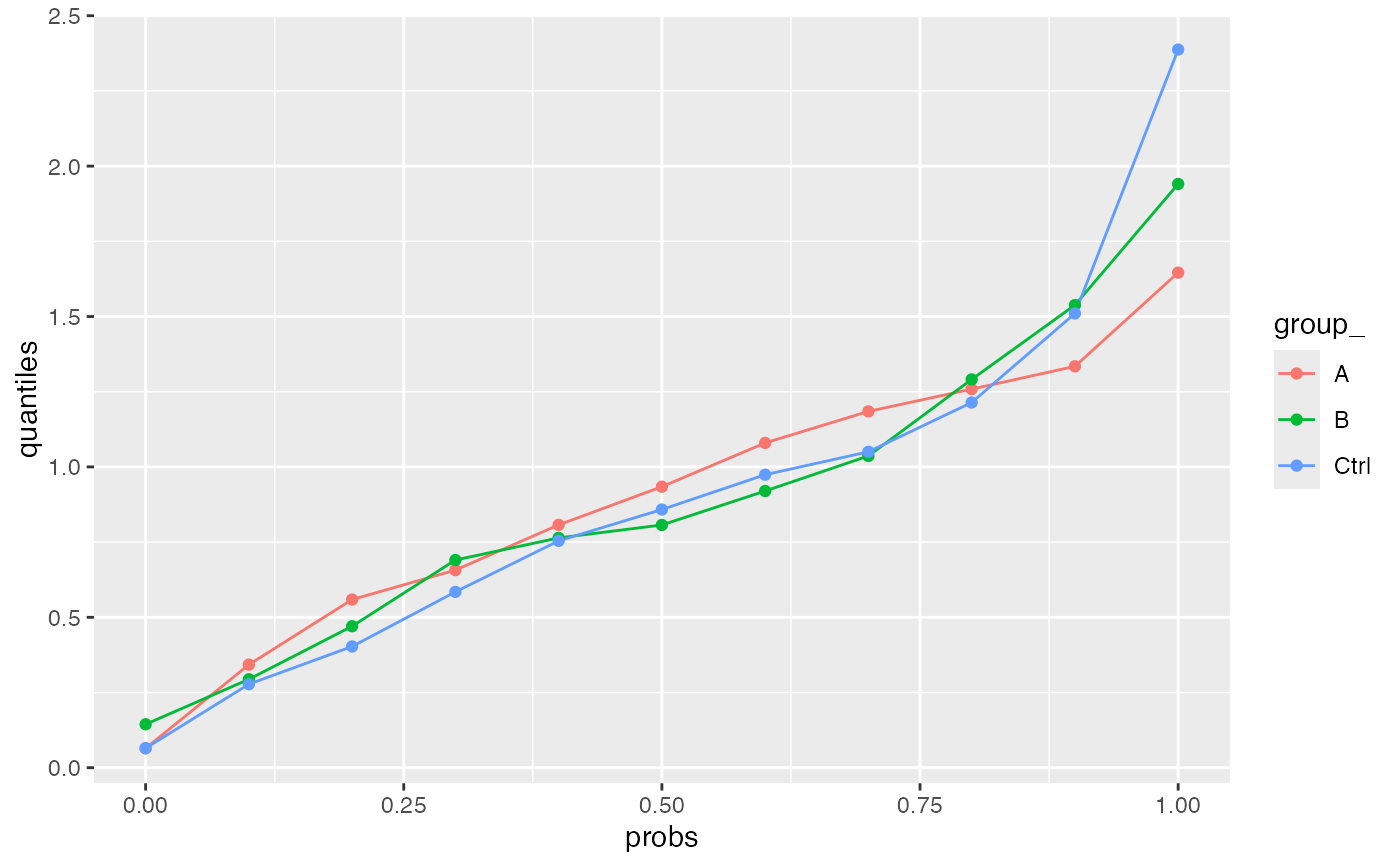
xx <- summarize_stats_quantiles(stats_res, lfq$relevant_factor_keys(), probs = seq(0, 1, by = 0.1))
ggplot2::ggplot(xx$long, aes(x = probs, y = quantiles, color = group_)) + geom_line() + geom_point()