p-value of protein from p.value of the median fold change peptide.
Source:R/tidyMS_moderation.R
get_p_values_pbeta.Rdp-value of protein from p.value of the median fold change peptide.
See also
Other modelling:
AnovaExtractor,
Contrasts,
ContrastsDEqMSFacade,
ContrastsDEqMSVoomFacade,
ContrastsFirth,
ContrastsFirthFacade,
ContrastsLMFacade,
ContrastsLMImputeFacade,
ContrastsLMMissingFacade,
ContrastsLimma,
ContrastsLimmaFacade,
ContrastsLimmaImputeFacade,
ContrastsLimmaVoomFacade,
ContrastsLimmaVoomImputeFacade,
ContrastsLimpaFacade,
ContrastsLmerFacade,
ContrastsMissing,
ContrastsModerated,
ContrastsModeratedDEqMS,
ContrastsPlotter,
ContrastsRLMFacade,
ContrastsROPECA,
ContrastsROPECAFacade,
ContrastsTable,
INTERNAL_FUNCTIONS_BY_FAMILY,
LR_test(),
Model,
ModelFirth,
ModelLimma,
StrategyLM,
StrategyLimma,
StrategyLimpa,
StrategyLmer,
StrategyLogistf,
StrategyRLM,
build_contrast_analysis(),
build_model(),
build_model_glm_peptide(),
build_model_glm_protein(),
build_model_impute(),
build_model_limma(),
build_model_limma_impute(),
build_model_limma_voom(),
build_model_limma_voom_impute(),
build_model_limpa(),
build_model_logistf(),
compute_borrowed_variance(),
compute_borrowed_variance_limma(),
compute_contrast(),
compute_lmer_contrast(),
contrasts_fisher_exact(),
get_anova_df(),
get_complete_model_fit(),
group_label(),
impute_refit_singular(),
is_singular_lm(),
linfct_all_possible_contrasts(),
linfct_factors_contrasts(),
linfct_from_model(),
linfct_matrix_contrasts(),
merge_contrasts_results(),
model_analyse(),
model_summary(),
moderated_p_deqms(),
moderated_p_deqms_long(),
moderated_p_limma(),
moderated_p_limma_long(),
new_lm_imputed(),
pivot_model_contrasts_to_wide(),
plot_lmer_peptide_predictions(),
sim_build_models_lm(),
sim_build_models_lmer(),
sim_build_models_logistf(),
sim_make_model_lm(),
sim_make_model_lmer(),
strategy_limma(),
strategy_limpa(),
strategy_logistf(),
summary_ROPECA_median_p.scaled()
Examples
plot(get_p_values_pbeta(0.1,1:10,10), ylim=c(0,0.1))
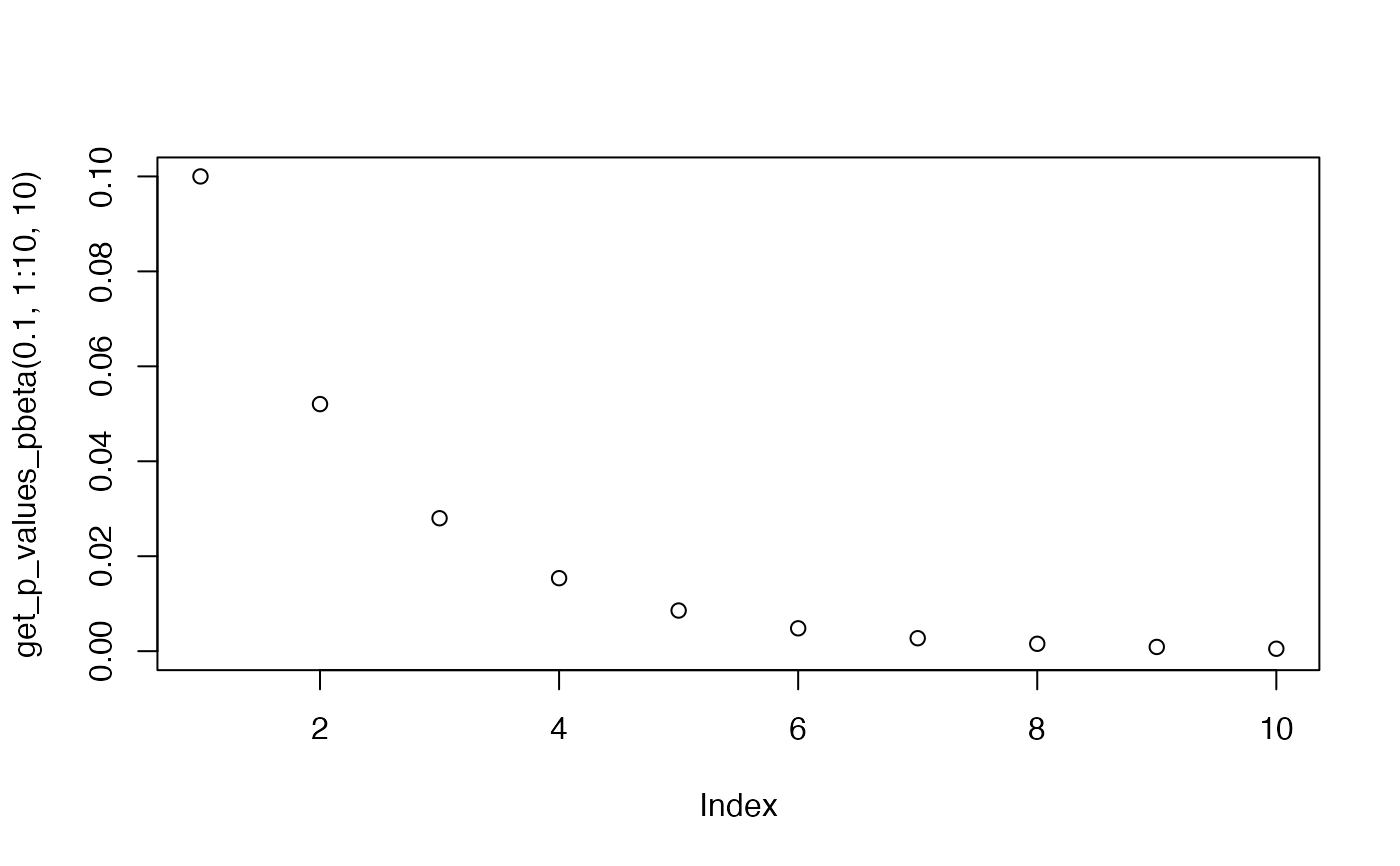 plot(get_p_values_pbeta(0.1,1:10,3), ylim=c(0,0.1))
plot(get_p_values_pbeta(0.1,1:10,3), ylim=c(0,0.1))
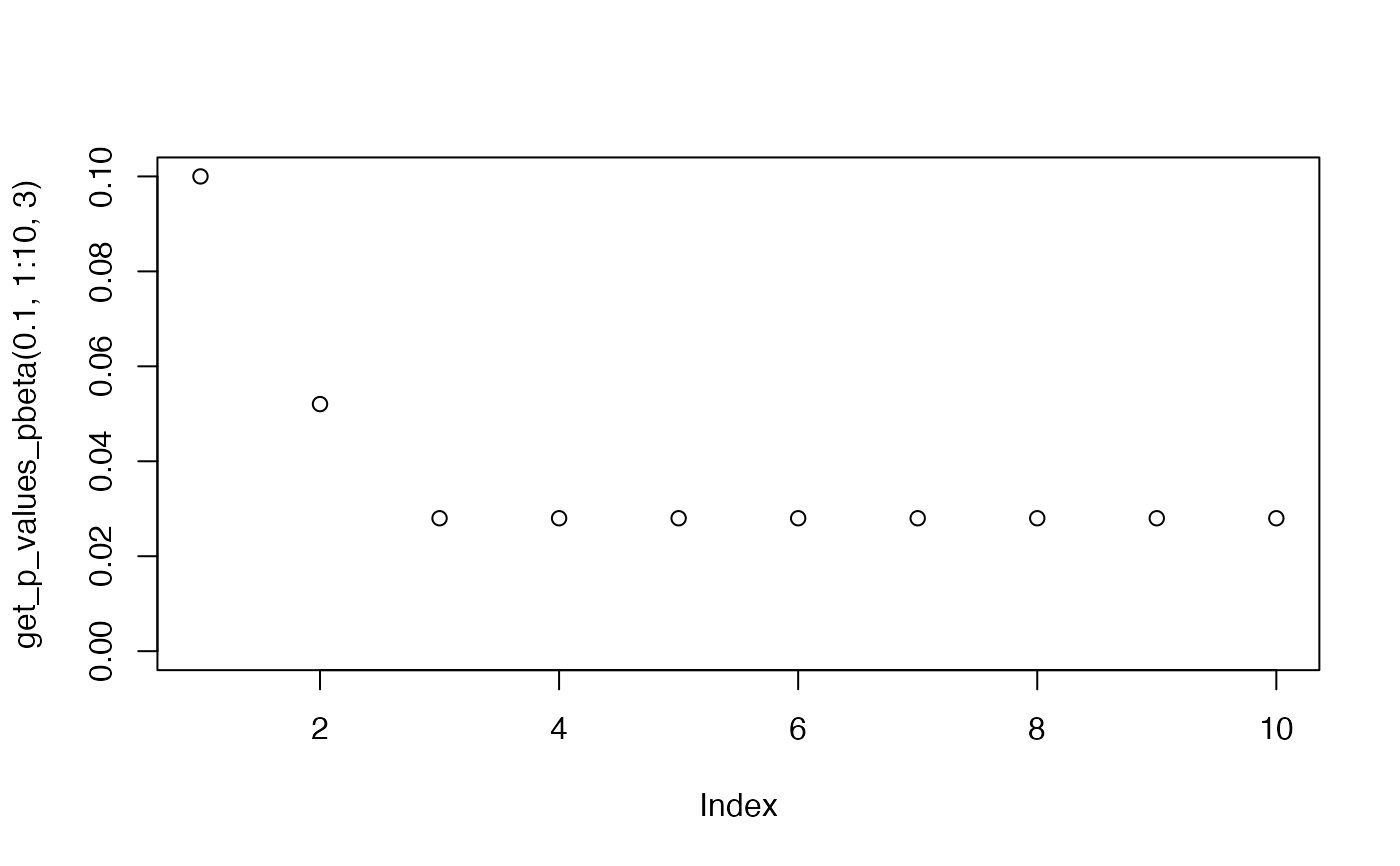 plot(get_p_values_pbeta(0.3,1:30, 3), ylim=c(0,0.1))
abline(h=.05,col = 2)
plot(get_p_values_pbeta(0.3,1:30, 3), ylim=c(0,0.1))
abline(h=.05,col = 2)
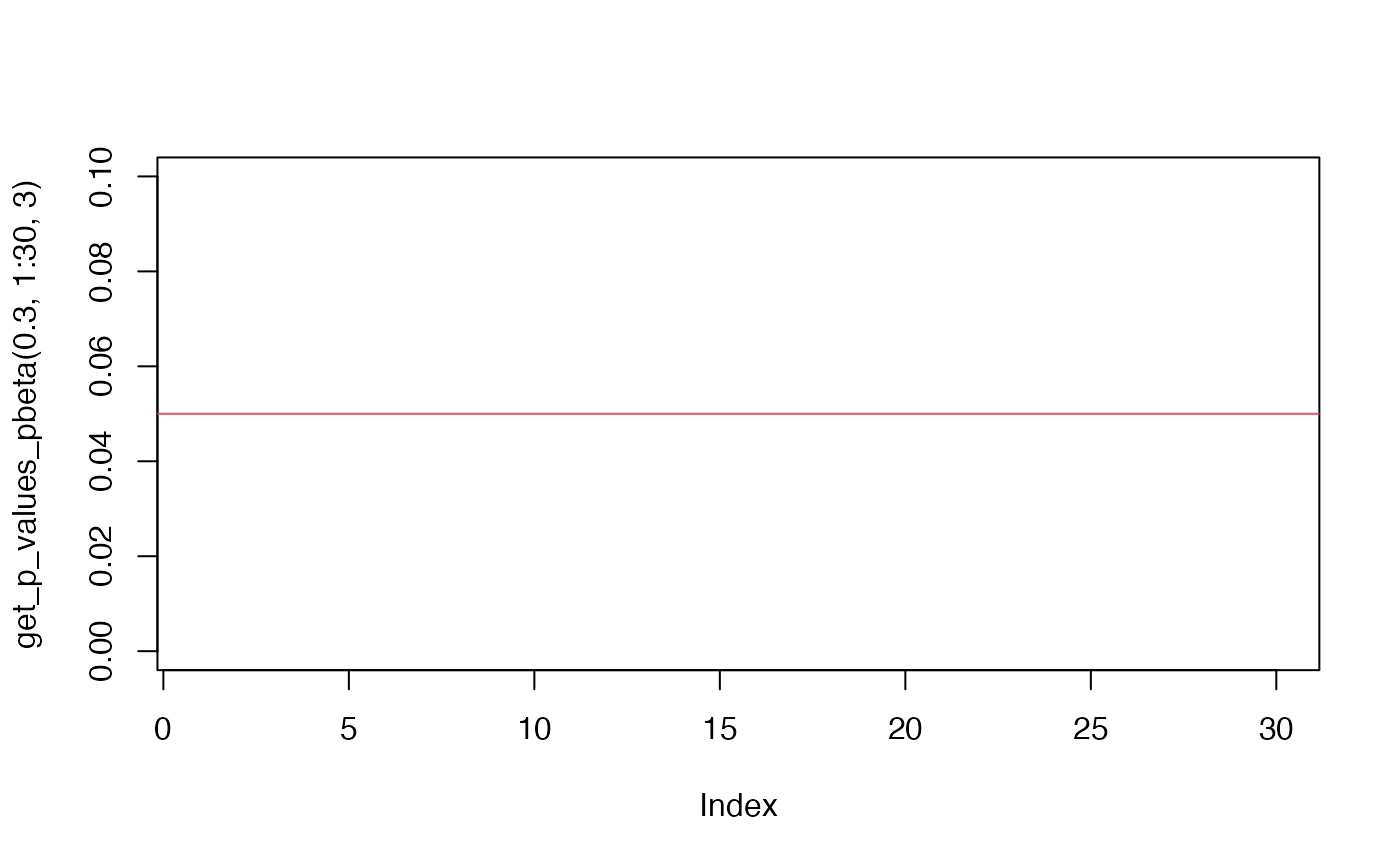 plot(seq(0.0,1.0,length=30),get_p_values_pbeta(seq(0.0,1.0,length=30),rep(10,30)))
abline(0,1)
plot(seq(0.0,1.0,length=30),get_p_values_pbeta(seq(0.0,1.0,length=30),rep(10,30)))
abline(0,1)
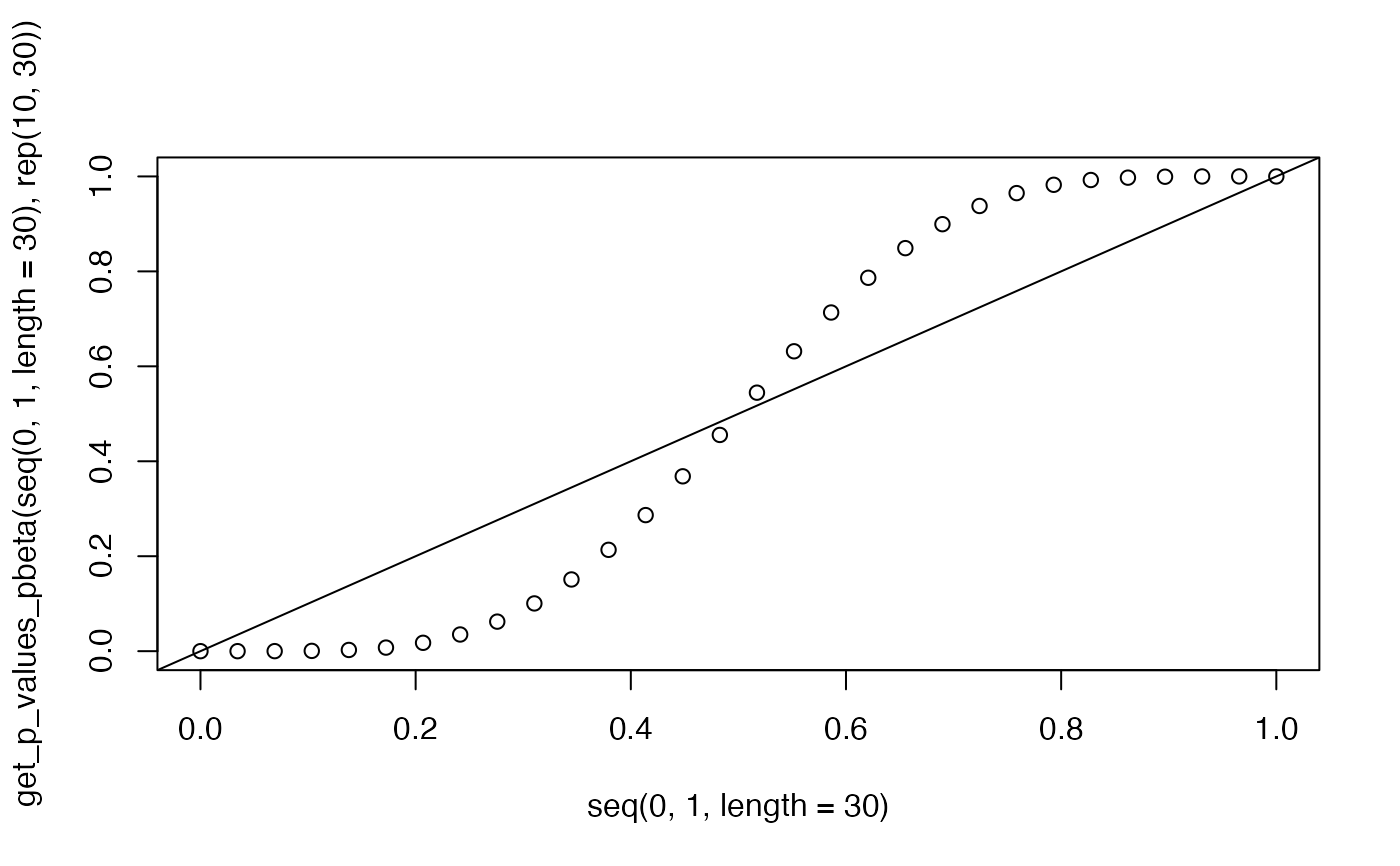 plot(seq(0.0,1.0,length=30),get_p_values_pbeta(seq(0.0,1.0,length=30),rep(10,30),3))
abline(0,1)
plot(seq(0.0,1.0,length=30),get_p_values_pbeta(seq(0.0,1.0,length=30),rep(10,30),3))
abline(0,1)
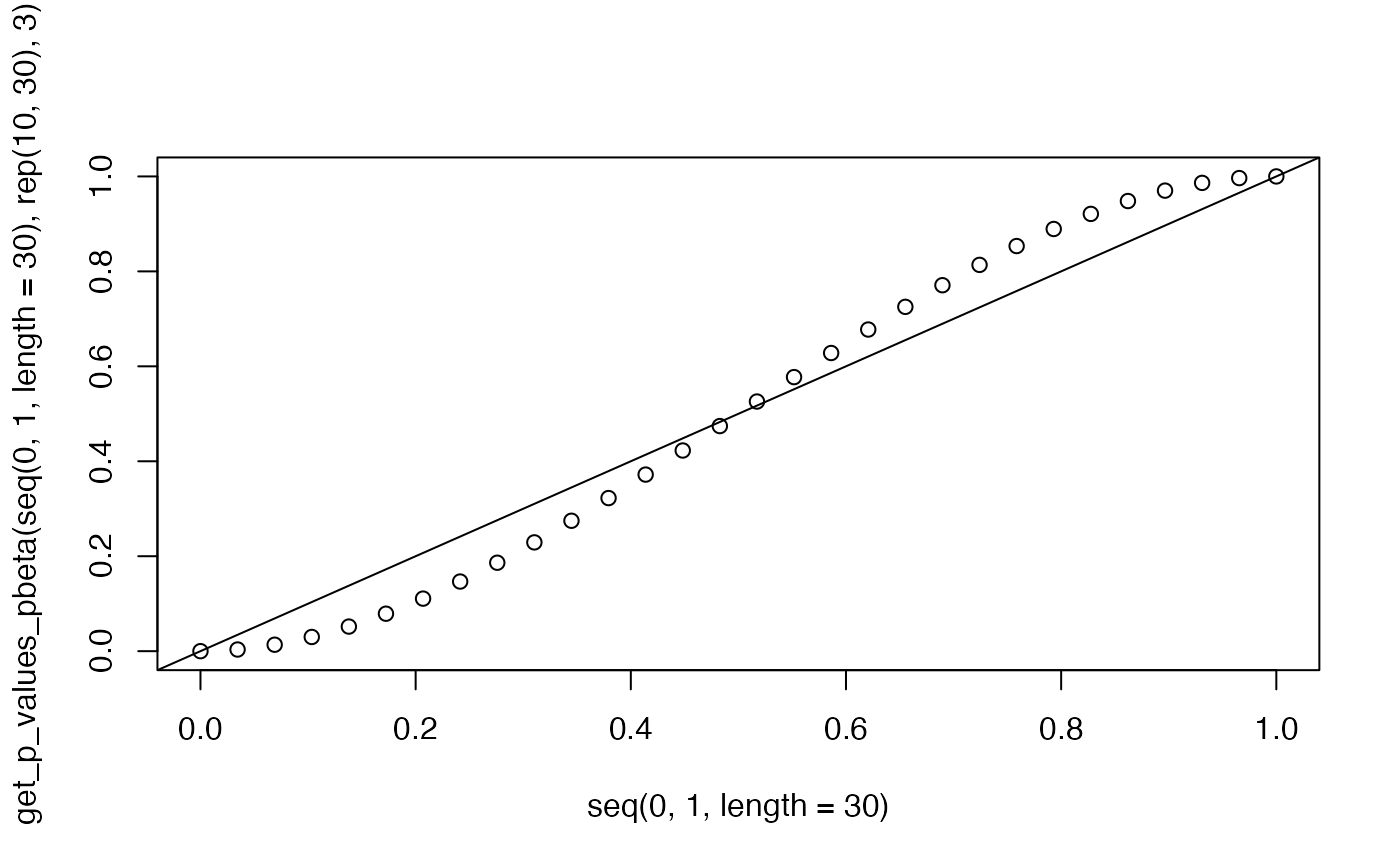 testthat::expect_equal(get_p_values_pbeta(0.3,10, 3),0.216, tolerance = 1e-4)
testthat::expect_equal(get_p_values_pbeta(0,10, 3),0, tolerance = 1e-4)
testthat::expect_equal(get_p_values_pbeta(1,10, 3),1, tolerance = 1e-4)
testthat::expect_equal(get_p_values_pbeta(1,10, 3),get_p_values_pbeta(1,3, 10), tolerance = 1e-4)
testthat::expect_equal(get_p_values_pbeta(0.3,10, 3),0.216, tolerance = 1e-4)
testthat::expect_equal(get_p_values_pbeta(0,10, 3),0, tolerance = 1e-4)
testthat::expect_equal(get_p_values_pbeta(1,10, 3),1, tolerance = 1e-4)
testthat::expect_equal(get_p_values_pbeta(1,10, 3),get_p_values_pbeta(1,3, 10), tolerance = 1e-4)