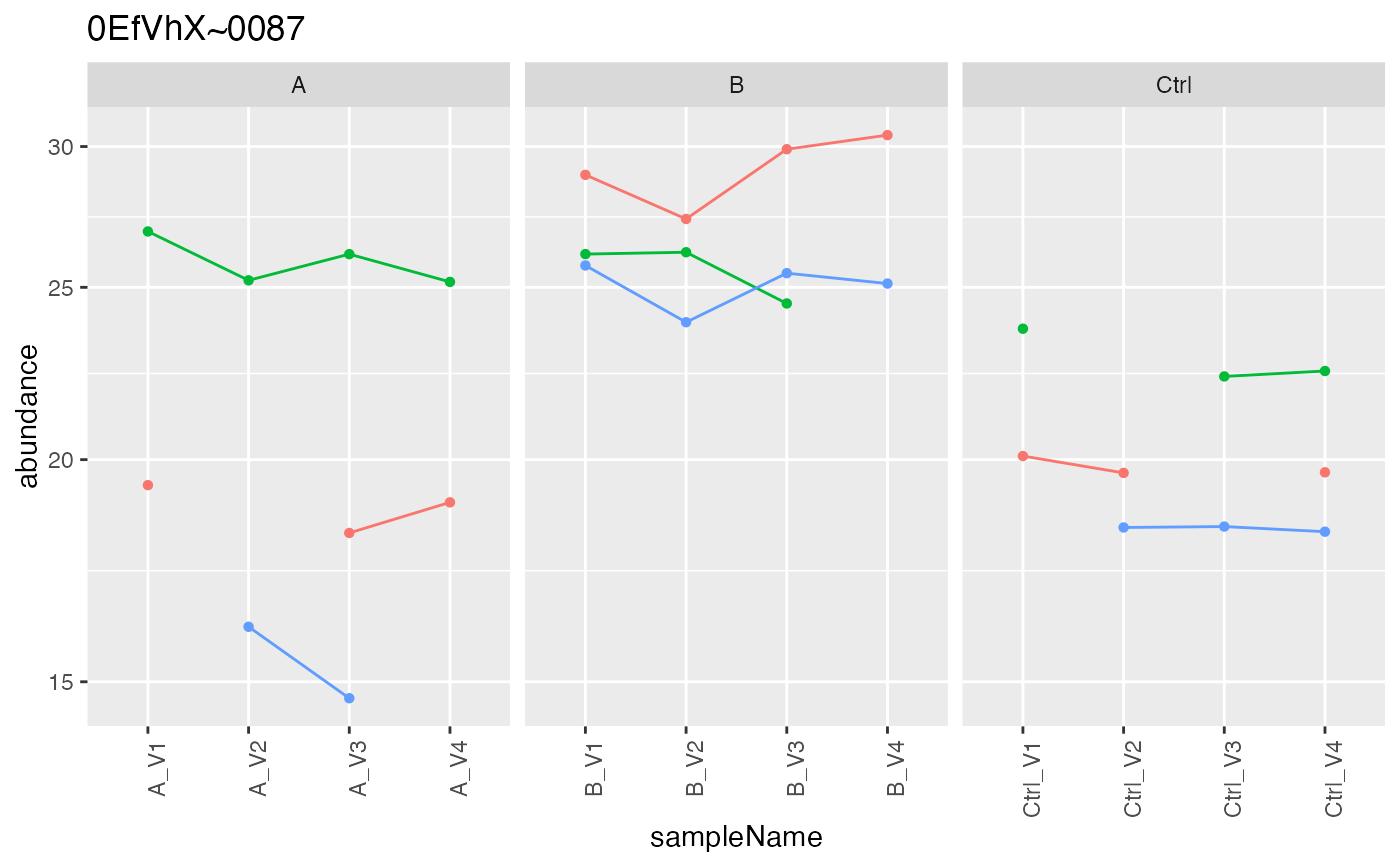
Plot peptide intensities of protein as a function of the sample and factor
Source:R/tidyMS_aggregation.R
plot_hierarchies_line.RdPlot peptide intensities of protein as a function of the sample and factor
See also
Other aggregation:
INTERNAL_FUNCTIONS_BY_FAMILY,
aggregate_intensity_top_n(),
estimate_intensity(),
medpolish_estimate(),
medpolish_estimate_df(),
medpolish_estimate_dfconfig(),
plot_estimate(),
plot_hierarchies_add_quantline(),
plot_hierarchies_line_df(),
rlm_estimate(),
rlm_estimate_dfconfig()
Other plotting:
ContrastsPlotter,
INTERNAL_FUNCTIONS_BY_FAMILY,
medpolish_estimate_df(),
missigness_histogram(),
missingness_per_condition(),
missingness_per_condition_cumsum(),
plot_estimate(),
plot_heatmap(),
plot_heatmap_cor(),
plot_hierarchies_add_quantline(),
plot_hierarchies_boxplot_df(),
plot_hierarchies_line_df(),
plot_intensity_distribution_violin(),
plot_na_heatmap(),
plot_pca(),
plot_raster(),
plot_sample_correlation(),
upset_interaction_missing_stats(),
upset_missing_stats()
Examples
istar <- sim_lfq_data_peptide_config()
#> creating sampleName from file_name column
#> completing cases
#> completing cases done
#> setup done
config <- istar$config
analysis <- istar$data
xnested <- analysis |>
dplyr::group_by(across(all_of(config$hierarchy_keys_depth()))) |>
tidyr::nest()
lfq <- LFQData$new(analysis, config)
prolfqua::plot_hierarchies_line(xnested$data[[1]], xnested$protein_Id[[1]], lfq)
#> Warning: Removed 7 rows containing missing values or values outside the scale range
#> (`geom_point()`).
#> Warning: Removed 4 rows containing missing values or values outside the scale range
#> (`geom_line()`).