R6 class representing modelling result
R6 class representing modelling result
See also
Other modelling:
AnovaExtractor,
Contrasts,
ContrastsDEqMSFacade,
ContrastsDEqMSVoomFacade,
ContrastsFirth,
ContrastsFirthFacade,
ContrastsLMFacade,
ContrastsLMImputeFacade,
ContrastsLMMissingFacade,
ContrastsLimma,
ContrastsLimmaFacade,
ContrastsLimmaImputeFacade,
ContrastsLimmaVoomFacade,
ContrastsLimmaVoomImputeFacade,
ContrastsLimpaFacade,
ContrastsLmerFacade,
ContrastsMissing,
ContrastsModerated,
ContrastsModeratedDEqMS,
ContrastsPlotter,
ContrastsRLMFacade,
ContrastsROPECA,
ContrastsROPECAFacade,
ContrastsTable,
INTERNAL_FUNCTIONS_BY_FAMILY,
LR_test(),
Model,
ModelLimma,
StrategyLM,
StrategyLimma,
StrategyLimpa,
StrategyLmer,
StrategyLogistf,
StrategyRLM,
build_contrast_analysis(),
build_model(),
build_model_glm_peptide(),
build_model_glm_protein(),
build_model_impute(),
build_model_limma(),
build_model_limma_impute(),
build_model_limma_voom(),
build_model_limma_voom_impute(),
build_model_limpa(),
build_model_logistf(),
compute_borrowed_variance(),
compute_borrowed_variance_limma(),
compute_contrast(),
compute_lmer_contrast(),
contrasts_fisher_exact(),
get_anova_df(),
get_complete_model_fit(),
get_p_values_pbeta(),
group_label(),
impute_refit_singular(),
is_singular_lm(),
linfct_all_possible_contrasts(),
linfct_factors_contrasts(),
linfct_from_model(),
linfct_matrix_contrasts(),
merge_contrasts_results(),
model_analyse(),
model_summary(),
moderated_p_deqms(),
moderated_p_deqms_long(),
moderated_p_limma(),
moderated_p_limma_long(),
new_lm_imputed(),
pivot_model_contrasts_to_wide(),
plot_lmer_peptide_predictions(),
sim_build_models_lm(),
sim_build_models_lmer(),
sim_build_models_logistf(),
sim_make_model_lm(),
sim_make_model_lmer(),
strategy_limma(),
strategy_limpa(),
strategy_logistf(),
summary_ROPECA_median_p.scaled()
Super class
prolfqua::ModelInterface -> ModelFirth
Public fields
modelsdata.frame with modelling data and model.
model_namename of model
subject_ide.g. protein_Id
anova_dffunction to compute anova
p.adjustfunction to adjust p-values
Methods
Method new()
initialize
Usage
ModelFirth$new(
models,
model_name = "modelFirth",
subject_id = "protein_Id",
p.adjust = prolfqua::adjust_p_values
)Method anova_histogram()
histogram of ANOVA results
Usage
ModelFirth$anova_histogram(what = c("p.value", "FDR"))Examples
istar <- prolfqua::sim_lfq_data_peptide_config(Nprot = 10, with_missing = TRUE,
weight_missing = 0.5, seed = 3)
#> creating sampleName from file_name column
#> completing cases
#> completing cases done
#> setup done
istar$data <- prolfqua::encode_bin_resp(LFQData$new(istar$data, istar$config))
istar$config$bin_resp <- "bin_resp"
tmp <- LFQData$new(istar$data, istar$config)
formula <- paste0(tmp$get_config()$bin_resp , "~ group_")
mod <- build_model_logistf(tmp, formula)
#> Joining with `by = join_by(protein_Id)`
#> Joining with `by = join_by(protein_Id)`
tmp <- mod$get_coefficients()
mod$coef_histogram()
#> $plot
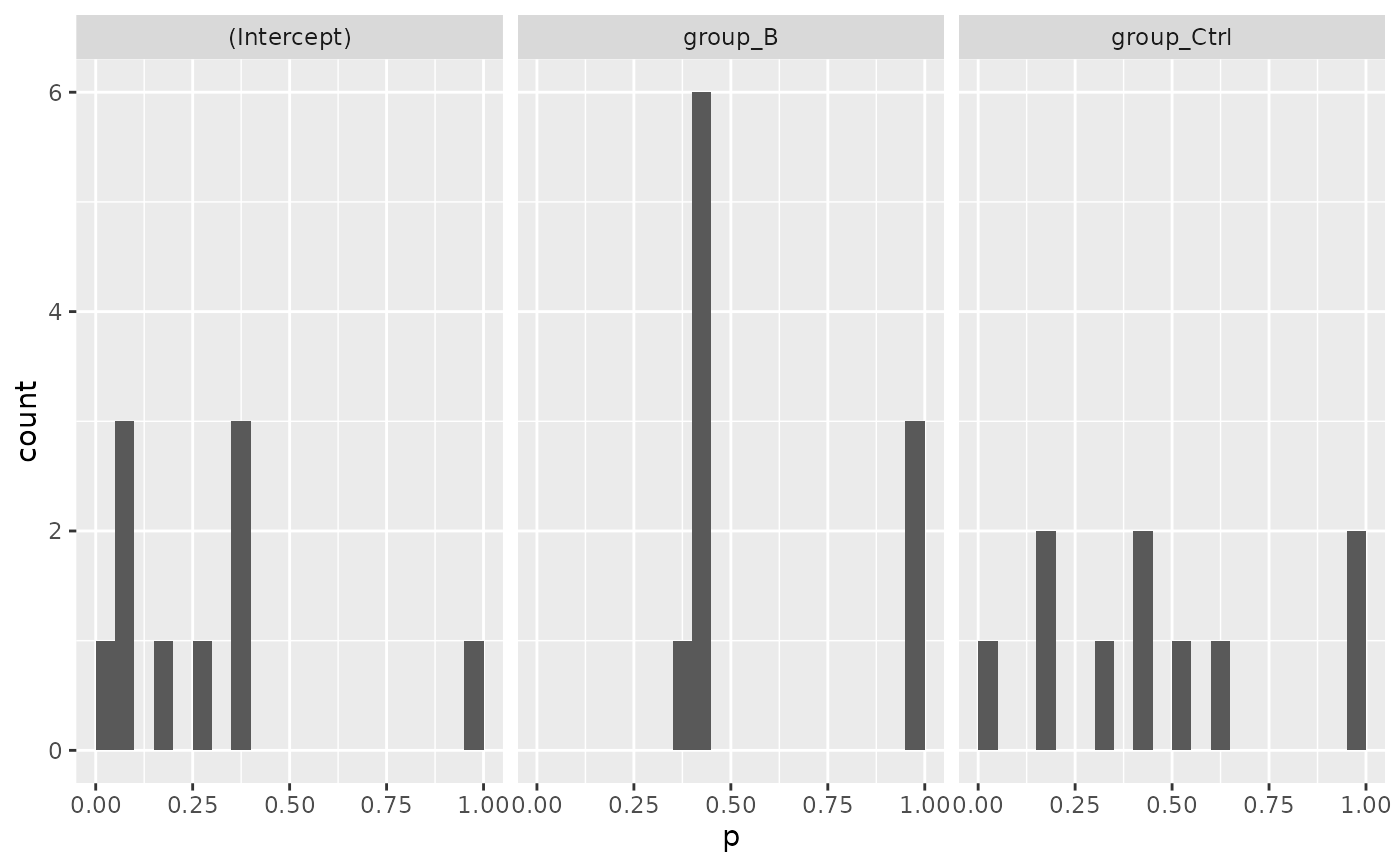 #>
#> $name
#> [1] "Coef_Histogram_modelFirth.pdf"
#>
mod$coef_pairs()
#> $plot
#> # A tibble: 10 × 4
#> subject_id `(Intercept)` group_B group_Ctrl
#> <chr> <dbl> <dbl> <dbl>
#> 1 7IZdVV~0841 2.10 6.77e- 1 -6.66e- 1
#> 2 AZPG26~2091 0.905 2.23e-16 1.82e+ 0
#> 3 AoNKbb~6908 1.01 -7.77e- 1 -4.25e- 1
#> 4 JnEDsz~4938 0.00838 8.36e-11 9.37e- 1
#> 5 tCZCHm~6695 2.16 1.23e+ 0 1.23e+ 0
#> 6 EnbuYk~0350 0.847 1.35e+ 0 -3.04e+ 0
#> 7 NNASGK~1417 2.20 -1.35e+ 0 4.13e-16
#> 8 XYrp6h~5793 2.20 -1.52e-15 -1.07e-15
#> 9 quTD7H~4566 0.847 1.35e+ 0 -8.47e- 1
#> 10 tHE075~8231 0.847 1.35e+ 0 1.35e+ 0
#>
#> $name
#> [1] "Coef_Pairsplot_modelFirth.pdf"
#>
mod$get_anova()
#> Warning: method not implemented!
#> NULL
mod$coef_volcano()
#> $plot
#>
#> $name
#> [1] "Coef_Histogram_modelFirth.pdf"
#>
mod$coef_pairs()
#> $plot
#> # A tibble: 10 × 4
#> subject_id `(Intercept)` group_B group_Ctrl
#> <chr> <dbl> <dbl> <dbl>
#> 1 7IZdVV~0841 2.10 6.77e- 1 -6.66e- 1
#> 2 AZPG26~2091 0.905 2.23e-16 1.82e+ 0
#> 3 AoNKbb~6908 1.01 -7.77e- 1 -4.25e- 1
#> 4 JnEDsz~4938 0.00838 8.36e-11 9.37e- 1
#> 5 tCZCHm~6695 2.16 1.23e+ 0 1.23e+ 0
#> 6 EnbuYk~0350 0.847 1.35e+ 0 -3.04e+ 0
#> 7 NNASGK~1417 2.20 -1.35e+ 0 4.13e-16
#> 8 XYrp6h~5793 2.20 -1.52e-15 -1.07e-15
#> 9 quTD7H~4566 0.847 1.35e+ 0 -8.47e- 1
#> 10 tHE075~8231 0.847 1.35e+ 0 1.35e+ 0
#>
#> $name
#> [1] "Coef_Pairsplot_modelFirth.pdf"
#>
mod$get_anova()
#> Warning: method not implemented!
#> NULL
mod$coef_volcano()
#> $plot
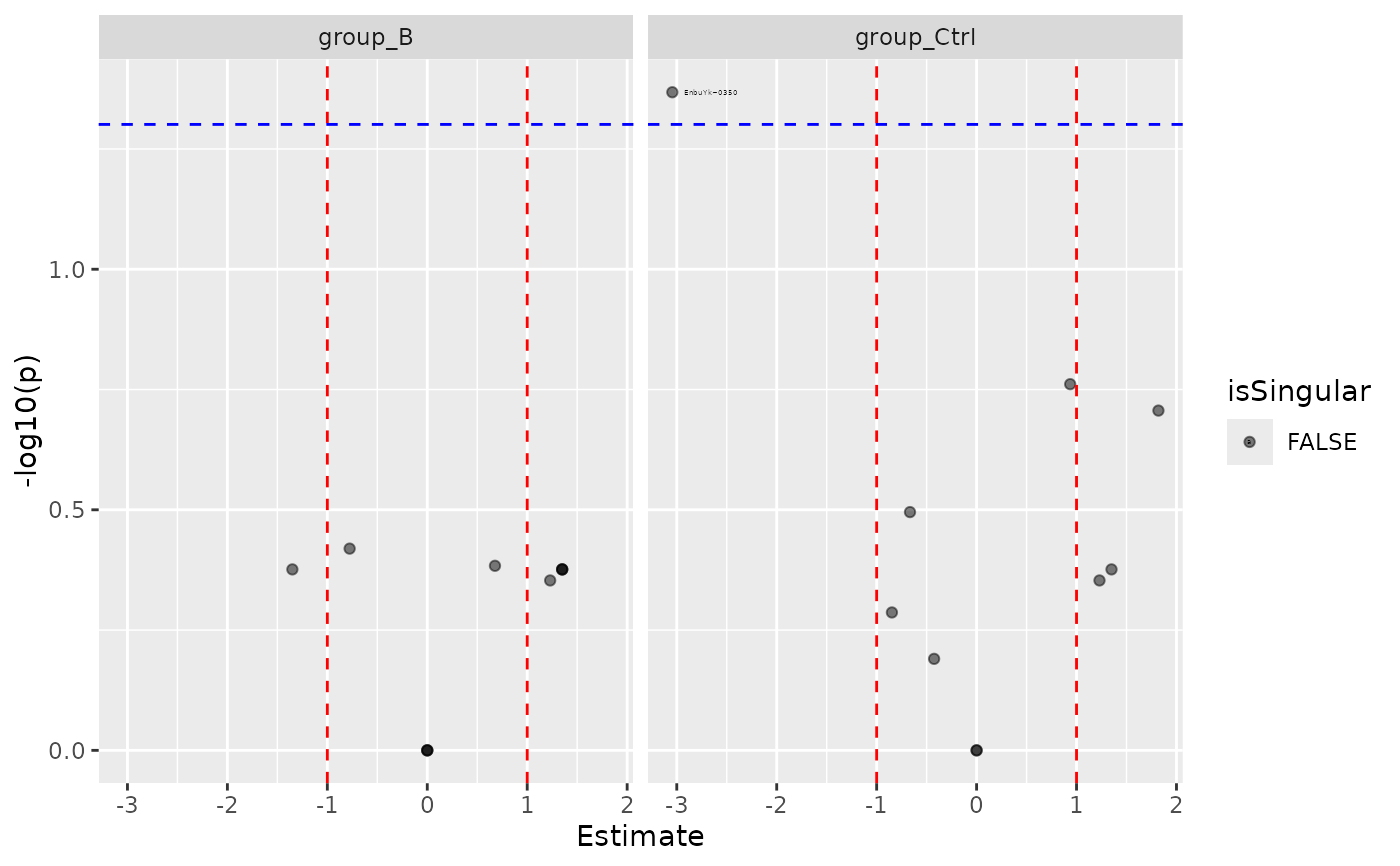 #>
#> $name
#> [1] "Coef_volcano_plot_modelFirth.pdf"
#>
mod$anova_histogram()
#> Warning: not implemented
#> NULL
mod$write_coef_figures(tempdir())
#> Writing figure into : /tmp/RtmpEpgSPK/Coef_Histogram_modelFirth.pdf
#> Writing figure into : /tmp/RtmpEpgSPK/Coef_volcano_plot_modelFirth.pdf
#> Writing figure into : /tmp/RtmpEpgSPK/Coef_Pairsplot_modelFirth.pdf
#> # A tibble: 10 × 4
#> subject_id `(Intercept)` group_B group_Ctrl
#> <chr> <dbl> <dbl> <dbl>
#> 1 7IZdVV~0841 2.10 6.77e- 1 -6.66e- 1
#> 2 AZPG26~2091 0.905 2.23e-16 1.82e+ 0
#> 3 AoNKbb~6908 1.01 -7.77e- 1 -4.25e- 1
#> 4 JnEDsz~4938 0.00838 8.36e-11 9.37e- 1
#> 5 tCZCHm~6695 2.16 1.23e+ 0 1.23e+ 0
#> 6 EnbuYk~0350 0.847 1.35e+ 0 -3.04e+ 0
#> 7 NNASGK~1417 2.20 -1.35e+ 0 4.13e-16
#> 8 XYrp6h~5793 2.20 -1.52e-15 -1.07e-15
#> 9 quTD7H~4566 0.847 1.35e+ 0 -8.47e- 1
#> 10 tHE075~8231 0.847 1.35e+ 0 1.35e+ 0
#> agg_record_8e495c993621
#> 2
istar <- prolfqua::sim_lfq_data_protein_config(Nprot = 10, with_missing = TRUE,
weight_missing = 0.5, seed = 3)
#> creating sampleName from file_name column
#> completing cases
#> completing cases done
#> setup done
istar$data <- prolfqua::encode_bin_resp(LFQData$new(istar$data, istar$config))
istar$config$bin_resp <- "bin_resp"
tmp <- LFQData$new(istar$data, istar$config)
formula <- paste0(tmp$get_config()$bin_resp , "~ group_")
mod <- build_model_logistf(tmp, formula)
#> Joining with `by = join_by(protein_Id)`
tmp <- mod$get_coefficients()
stopifnot(nrow(tmp) == 30)
mod$coef_histogram()
#> $plot
#>
#> $name
#> [1] "Coef_volcano_plot_modelFirth.pdf"
#>
mod$anova_histogram()
#> Warning: not implemented
#> NULL
mod$write_coef_figures(tempdir())
#> Writing figure into : /tmp/RtmpEpgSPK/Coef_Histogram_modelFirth.pdf
#> Writing figure into : /tmp/RtmpEpgSPK/Coef_volcano_plot_modelFirth.pdf
#> Writing figure into : /tmp/RtmpEpgSPK/Coef_Pairsplot_modelFirth.pdf
#> # A tibble: 10 × 4
#> subject_id `(Intercept)` group_B group_Ctrl
#> <chr> <dbl> <dbl> <dbl>
#> 1 7IZdVV~0841 2.10 6.77e- 1 -6.66e- 1
#> 2 AZPG26~2091 0.905 2.23e-16 1.82e+ 0
#> 3 AoNKbb~6908 1.01 -7.77e- 1 -4.25e- 1
#> 4 JnEDsz~4938 0.00838 8.36e-11 9.37e- 1
#> 5 tCZCHm~6695 2.16 1.23e+ 0 1.23e+ 0
#> 6 EnbuYk~0350 0.847 1.35e+ 0 -3.04e+ 0
#> 7 NNASGK~1417 2.20 -1.35e+ 0 4.13e-16
#> 8 XYrp6h~5793 2.20 -1.52e-15 -1.07e-15
#> 9 quTD7H~4566 0.847 1.35e+ 0 -8.47e- 1
#> 10 tHE075~8231 0.847 1.35e+ 0 1.35e+ 0
#> agg_record_8e495c993621
#> 2
istar <- prolfqua::sim_lfq_data_protein_config(Nprot = 10, with_missing = TRUE,
weight_missing = 0.5, seed = 3)
#> creating sampleName from file_name column
#> completing cases
#> completing cases done
#> setup done
istar$data <- prolfqua::encode_bin_resp(LFQData$new(istar$data, istar$config))
istar$config$bin_resp <- "bin_resp"
tmp <- LFQData$new(istar$data, istar$config)
formula <- paste0(tmp$get_config()$bin_resp , "~ group_")
mod <- build_model_logistf(tmp, formula)
#> Joining with `by = join_by(protein_Id)`
tmp <- mod$get_coefficients()
stopifnot(nrow(tmp) == 30)
mod$coef_histogram()
#> $plot
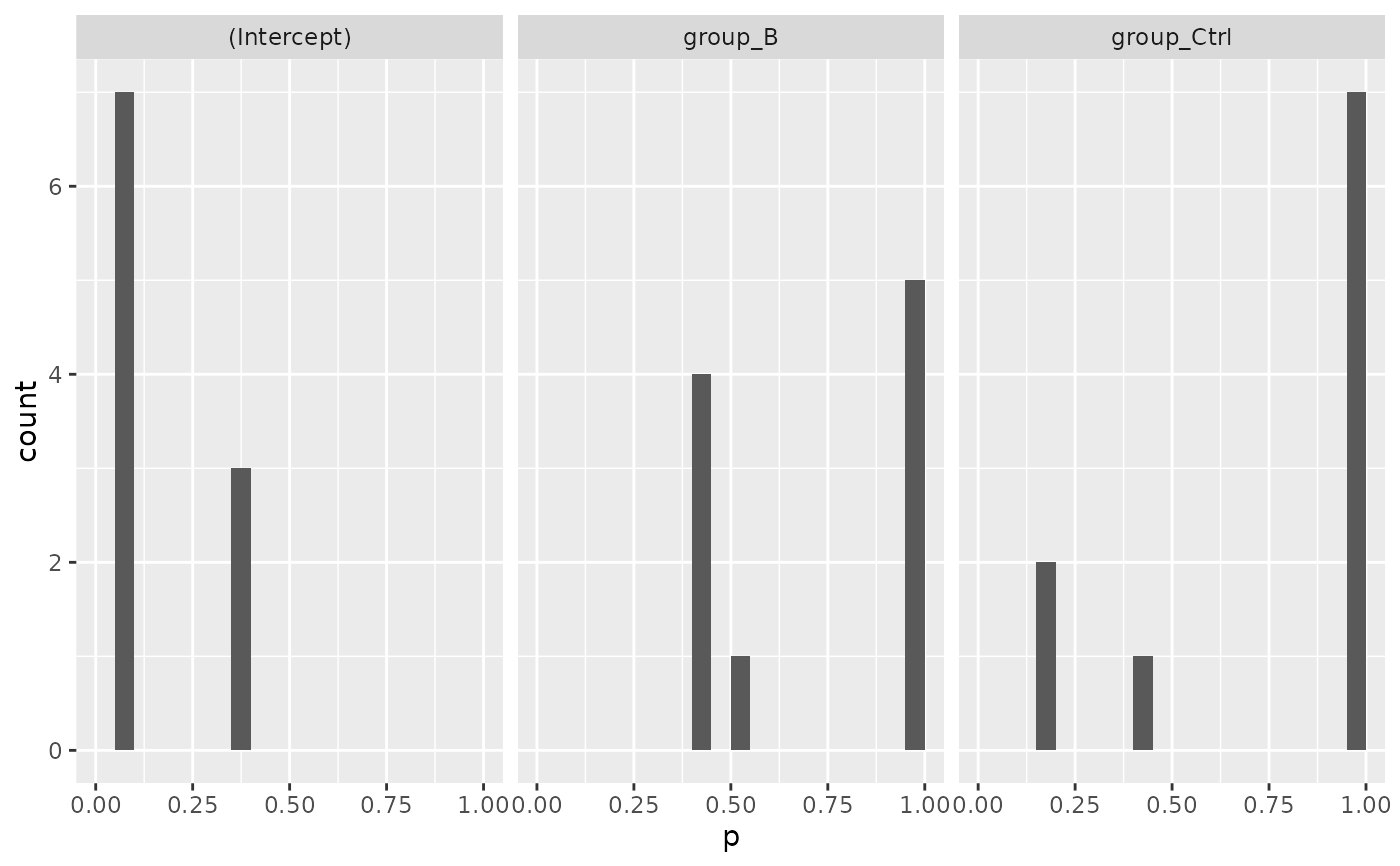 #>
#> $name
#> [1] "Coef_Histogram_modelFirth.pdf"
#>
mod$coef_pairs()
#> $plot
#> # A tibble: 10 × 4
#> subject_id `(Intercept)` group_B group_Ctrl
#> <chr> <dbl> <dbl> <dbl>
#> 1 7IZdVV~0841 2.20 5.61e-16 -2.20e+ 0
#> 2 AZPG26~2091 2.20 -1.52e-15 -1.07e-15
#> 3 AoNKbb~6908 2.20 6.16e-16 -1.35e+ 0
#> 4 EnbuYk~0350 0.847 -8.47e- 1 -6.83e-16
#> 5 JnEDsz~4938 -0.847 -1.35e+ 0 1.18e-16
#> 6 NNASGK~1417 2.20 -1.52e-15 -1.07e-15
#> 7 XYrp6h~5793 2.20 -1.52e-15 -1.07e-15
#> 8 quTD7H~4566 2.20 -1.35e+ 0 -2.20e+ 0
#> 9 tCZCHm~6695 2.20 -1.35e+ 0 4.13e-16
#> 10 tHE075~8231 0.847 1.35e+ 0 1.85e-16
#>
#> $name
#> [1] "Coef_Pairsplot_modelFirth.pdf"
#>
mod$get_anova()
#> Warning: method not implemented!
#> NULL
mod$coef_volcano()
#> $plot
#>
#> $name
#> [1] "Coef_Histogram_modelFirth.pdf"
#>
mod$coef_pairs()
#> $plot
#> # A tibble: 10 × 4
#> subject_id `(Intercept)` group_B group_Ctrl
#> <chr> <dbl> <dbl> <dbl>
#> 1 7IZdVV~0841 2.20 5.61e-16 -2.20e+ 0
#> 2 AZPG26~2091 2.20 -1.52e-15 -1.07e-15
#> 3 AoNKbb~6908 2.20 6.16e-16 -1.35e+ 0
#> 4 EnbuYk~0350 0.847 -8.47e- 1 -6.83e-16
#> 5 JnEDsz~4938 -0.847 -1.35e+ 0 1.18e-16
#> 6 NNASGK~1417 2.20 -1.52e-15 -1.07e-15
#> 7 XYrp6h~5793 2.20 -1.52e-15 -1.07e-15
#> 8 quTD7H~4566 2.20 -1.35e+ 0 -2.20e+ 0
#> 9 tCZCHm~6695 2.20 -1.35e+ 0 4.13e-16
#> 10 tHE075~8231 0.847 1.35e+ 0 1.85e-16
#>
#> $name
#> [1] "Coef_Pairsplot_modelFirth.pdf"
#>
mod$get_anova()
#> Warning: method not implemented!
#> NULL
mod$coef_volcano()
#> $plot
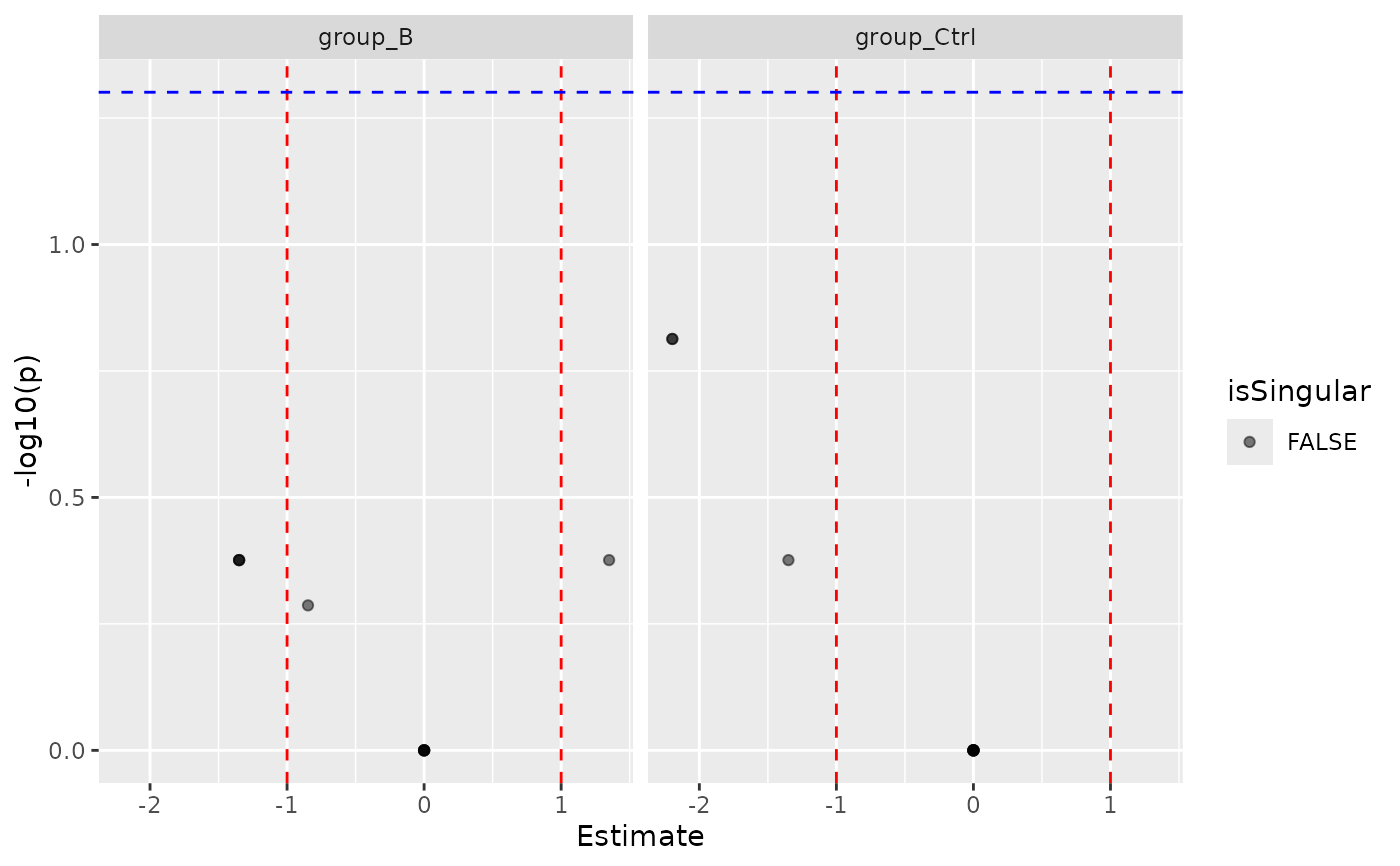 #>
#> $name
#> [1] "Coef_volcano_plot_modelFirth.pdf"
#>
mod$anova_histogram()
#> Warning: not implemented
#> NULL
mod$write_coef_figures(tempdir())
#> Writing figure into : /tmp/RtmpEpgSPK/Coef_Histogram_modelFirth.pdf
#> Writing figure into : /tmp/RtmpEpgSPK/Coef_volcano_plot_modelFirth.pdf
#> Writing figure into : /tmp/RtmpEpgSPK/Coef_Pairsplot_modelFirth.pdf
#> # A tibble: 10 × 4
#> subject_id `(Intercept)` group_B group_Ctrl
#> <chr> <dbl> <dbl> <dbl>
#> 1 7IZdVV~0841 2.20 5.61e-16 -2.20e+ 0
#> 2 AZPG26~2091 2.20 -1.52e-15 -1.07e-15
#> 3 AoNKbb~6908 2.20 6.16e-16 -1.35e+ 0
#> 4 EnbuYk~0350 0.847 -8.47e- 1 -6.83e-16
#> 5 JnEDsz~4938 -0.847 -1.35e+ 0 1.18e-16
#> 6 NNASGK~1417 2.20 -1.52e-15 -1.07e-15
#> 7 XYrp6h~5793 2.20 -1.52e-15 -1.07e-15
#> 8 quTD7H~4566 2.20 -1.35e+ 0 -2.20e+ 0
#> 9 tCZCHm~6695 2.20 -1.35e+ 0 4.13e-16
#> 10 tHE075~8231 0.847 1.35e+ 0 1.85e-16
#> agg_record_8e495c993621
#> 2
#>
#> $name
#> [1] "Coef_volcano_plot_modelFirth.pdf"
#>
mod$anova_histogram()
#> Warning: not implemented
#> NULL
mod$write_coef_figures(tempdir())
#> Writing figure into : /tmp/RtmpEpgSPK/Coef_Histogram_modelFirth.pdf
#> Writing figure into : /tmp/RtmpEpgSPK/Coef_volcano_plot_modelFirth.pdf
#> Writing figure into : /tmp/RtmpEpgSPK/Coef_Pairsplot_modelFirth.pdf
#> # A tibble: 10 × 4
#> subject_id `(Intercept)` group_B group_Ctrl
#> <chr> <dbl> <dbl> <dbl>
#> 1 7IZdVV~0841 2.20 5.61e-16 -2.20e+ 0
#> 2 AZPG26~2091 2.20 -1.52e-15 -1.07e-15
#> 3 AoNKbb~6908 2.20 6.16e-16 -1.35e+ 0
#> 4 EnbuYk~0350 0.847 -8.47e- 1 -6.83e-16
#> 5 JnEDsz~4938 -0.847 -1.35e+ 0 1.18e-16
#> 6 NNASGK~1417 2.20 -1.52e-15 -1.07e-15
#> 7 XYrp6h~5793 2.20 -1.52e-15 -1.07e-15
#> 8 quTD7H~4566 2.20 -1.35e+ 0 -2.20e+ 0
#> 9 tCZCHm~6695 2.20 -1.35e+ 0 4.13e-16
#> 10 tHE075~8231 0.847 1.35e+ 0 1.85e-16
#> agg_record_8e495c993621
#> 2